 samtools mpileup -uf reference.fasta file.bam bcftools call -c vcf2fq > sample.fq. These old versions remain available from the Sourceforge samtools project. I want to create consensus fasta sequence for long-read sequencing BAM files. However, assembly errors can occur due to data characteristics and assembly algorithms. Prior to the introduction of HTSlib, SAMtools and BCFtools were distributed Nanopore sequencing produces long reads and offers unique advantages over next-generation sequencing, especially for the assembly of draft bacterial genomes with improved completeness. Your specified prefix, so you may wish to add this directory to your $PATH: export PATH =/where/to/install/bin: $PATH # for sh or bash users setenv PATH /where/to/install/bin:$PATH # for csh users Historical SAMtools/BCFtools 0.1.x releases The executable programs will be installed to a bin subdirectory under See INSTALL in each of the source directories for further details. Building and installingīuilding each desired package from source is very simple: cd samtools-1.x # and similarly for bcftools and htslib New releases are announced on the samtools mailing lists and by Twitter. Or see the additional instructions in INSTALL to install them from a So you may also want to build and install HTSlib to get these utilities, HTSlib also provides the bgzip, htsfile, and tabix utilities, If you are writing your own programs against the HTSlib API. HTSlib is also distributed as a separate package which can be installed The code uses HTSlib internally, but these source packages contain their ownĬopies of htslib so they can be built independently. Sometimes there is the need to create a consensus sequence for an individual where the sequence incorporates variants typed for this individual. It regards an input file - as the standard input (stdin) and. How to count the number of mapped reads in a BAM or SAM file (SAM bitcode fields) more statistics about alignments. Load software modules module load samtools/1.11 htslib/1. The below steps are to filter and extract consensus gene sequences for each sample. Samtools is designed to work on a stream. get number of individual reads, paired reads that mapped both count double R1+R2. I joint call variants for 6 samples using FreeBayes (all.vcf), then bgzip and index it. It imports from and exports to the SAM (Sequence Alignment/Map) format, does sorting, merging and indexing, and allows one to retrieve reads in any regions swiftly. Writing list of indels 100 bases and longer to assm_indel_ge100.txt.Īll done, output written to assm_stats.txt, assm_summ.txt and assm_indel_ge100.txtĪll done, output written to assm_stats.txt, assm_summ.txt and assm_indel_ge100.SAMtools and BCFtools are distributed as individual packages. Samtools is a set of utilities that manipulate alignments in the BAM format. shape + ( 4 ,)) # set row colours: A blue, C red, G, green, T yello cols =. transpose () ppos = positions # create a figure p = figure ( title = "Base counts", plot_height = 300, plot_width = 600 ) img = np. # Plot feature array (click play) import aplanat from otting import figure from bokeh.models import Range1d # select just a region to plot reg = slice ( 3925, 3960 ) pdata = feature_array. Multiple chunks are returned if there are discontinuities in (pileup counts array, reference positions, insertion positions) :param weibull_summation: use a Weibull partial-counts approach, :param num_qstrat: number of layers for qscore stratification. :param keep_missing: whether to keep reads when tag is missing. :param tag_value: integer value of tag for reads to keep. :param tag_name: two letter tag name by which to filter reads. 
The results of this command / sequence legth 100 to have the genome covered. 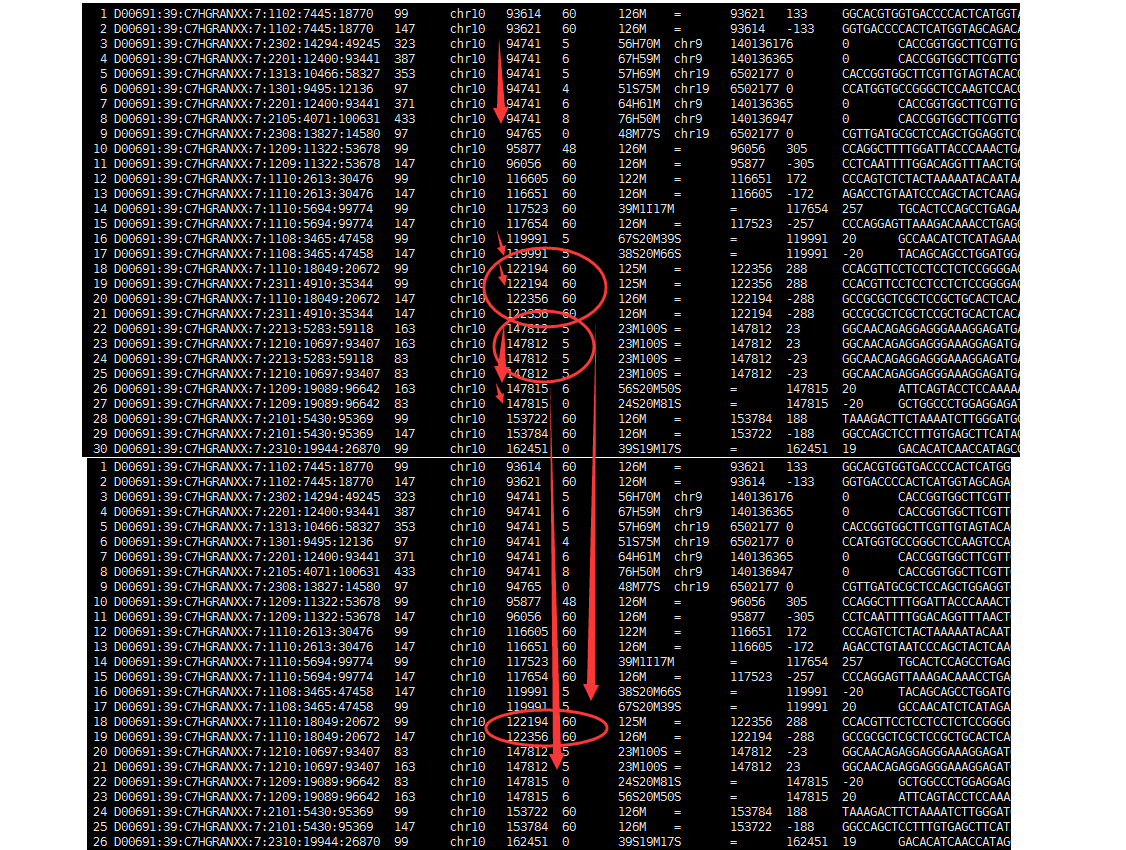 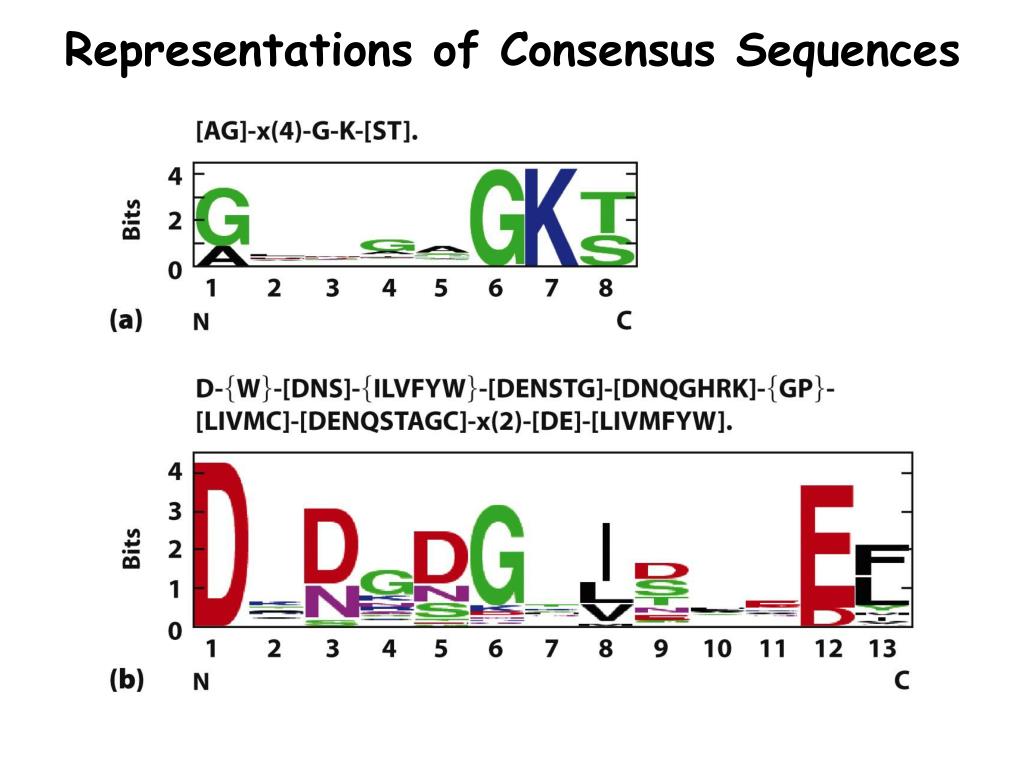
:param workers: worker threads for calculating pileup. samtools depth my.bam > qry-depth> wc -l qry-depth. Regions are processed in parallel and stitched before being returned. :param region_split: largest region to process in single thread. If `None` (or of length 1), counts are not split. If the FASTQ data has UMIs, it can be preprocessed using fastp to move the UMIs from read sequences to read identifiers.The main workflow of gencore is described in Fig. Overview As we have seen, the SAMTools suite allows you to manipulate the SAM/BAM files produced by most aligners. :param dtype_prefixes: prefixes for query names which to separate counts. gencore requires an input of position sorted BAM file and a reference genome FASTA file. Pileup_counts(region, bam, dtype_prefixes=None, region_split=100000, workers=8, tag_name=None, tag_value=None, keep_missing=False, num_qstrat=1, weibull_summation=False, read_group=None)Ĭreate pileup counts feature array for region. Help on function pileup_counts in module medaka.features: 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
